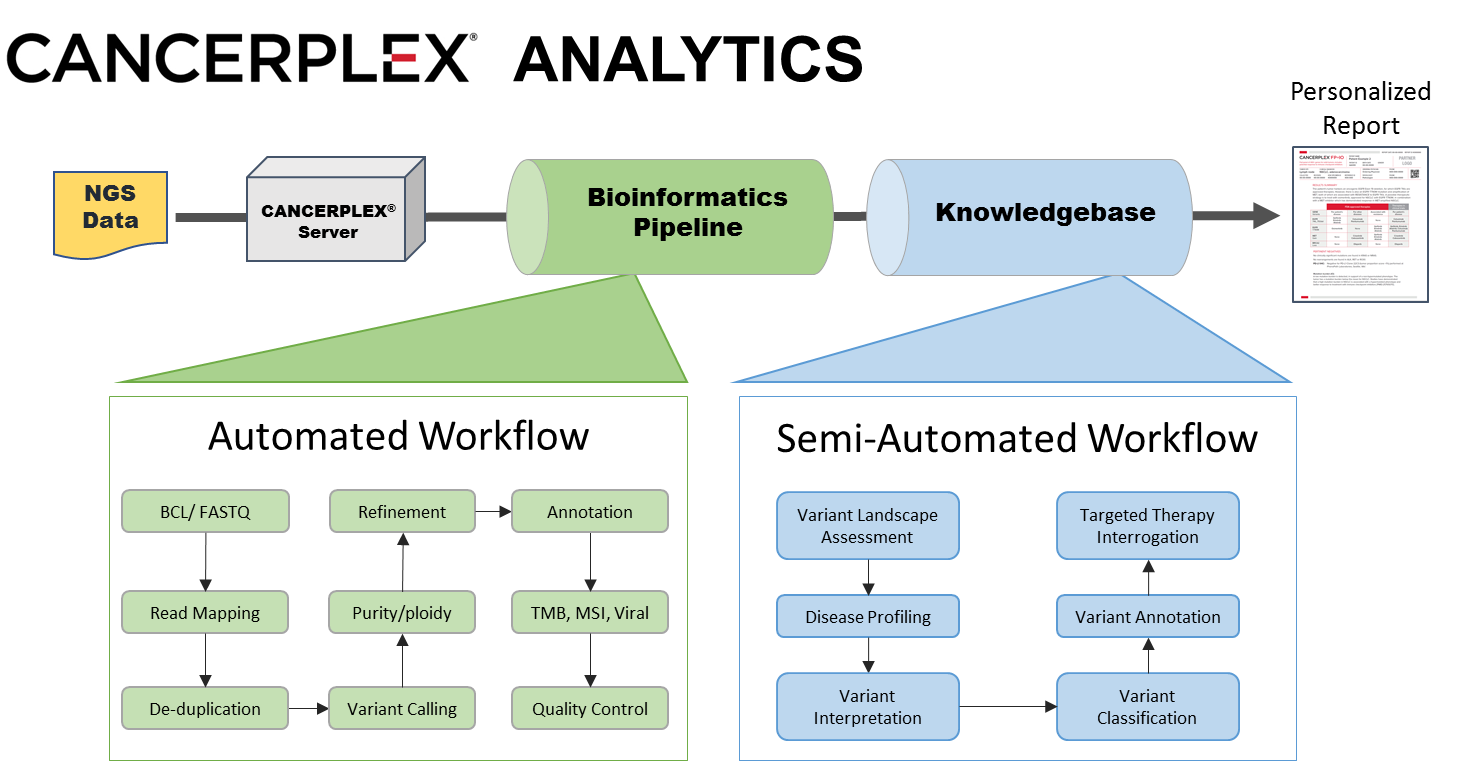

KEW BIOINFORMATICS
Comprehensive suite of state-of-art and proprietary algorithms for optimal and accurate genomic variant detection.
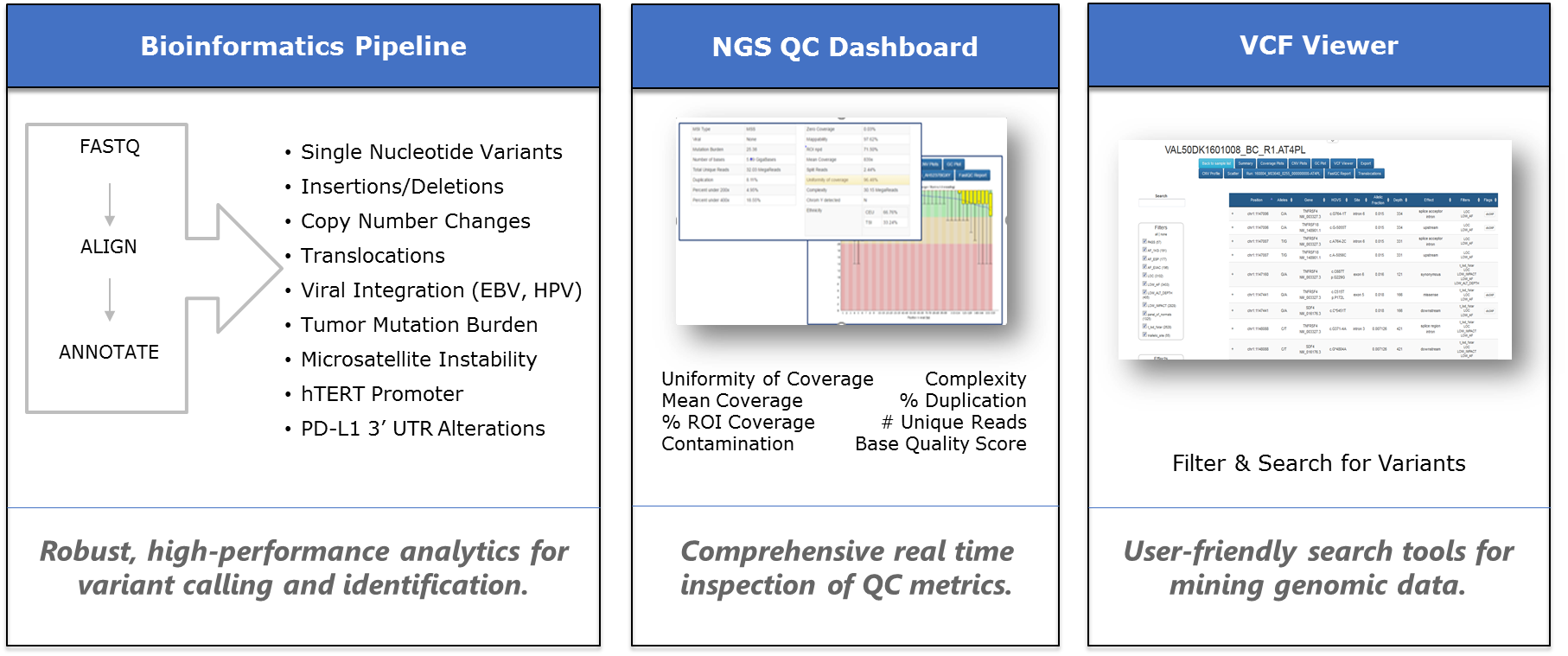

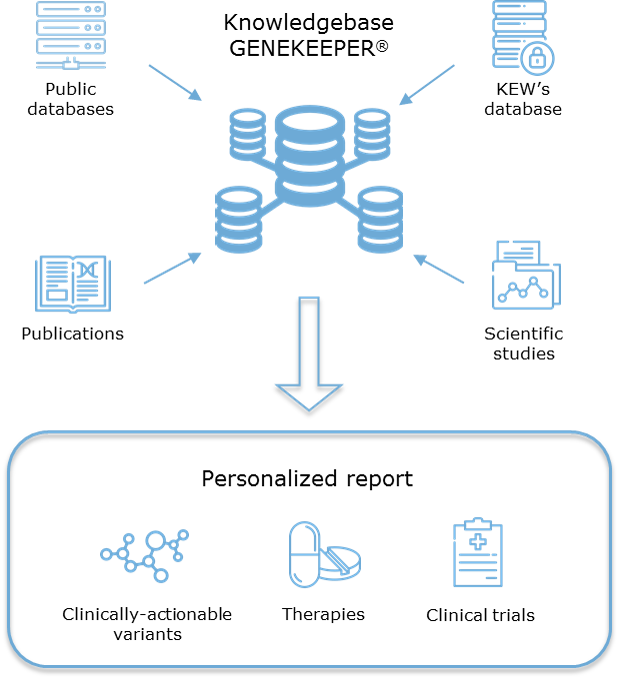
GENEKEEPER
GENEKEEPER® is a ‘State of the Art’ HIPAA-compliant knowledgebase and curation hub that facilitates analysis of patient mutations, amplifications, genomic alterations, and therapeutic response biomarkers in the most centralized high-throughput manner for timely and dependable delivery of patient reports.

BENEFITS of KEW’S ANALYTICS
Comprehensive Variant Annotation
KEW’s analytic platform and interpretation pipeline intersects data from its manually curated
GENEKEEPER knowledgebase, with real-time data feeds from 20+ industry leading protein, gene, variant, drug, and clinical trial databases for comprehensive curator-guided interpretation of variants for their impact on patient care.

AMP Guidelines
GENEKEEPER is one of the few databases that aligns with AMP, ASCO and CAP consensus guidelines [PMID:27993330] for interpretation and reporting of somatic variants in Next-Generation Sequencing–Based Oncology Panels. This standardized reporting methodology facilitates evidence-based variant categorization, annotation, and reporting in cancer genomics.
Scalability and Flexibility
GENEKEEPER is architected to accommodate varying levels of access for partners and research collaborators, and can be localized for languages other than English. Country/region-specific reports can be generated to provide customized drug approval data and clinical trial information.
Database Management
As the Cancer Biology and Personalized Medicine fields continue to evolve, GENEKEEPER functionalities and resources are expanded daily with high-quality expert-reviewed data, while in parallel ensuring its utility as a cutting-edge curation, data interpretation, and reporting hub.

Team of experts
Precision Medicine Scientists at KEW, all of which are PhD/MD trained experts in the fields of cancer biology, molecular biology, and oncology, carry out personalized analysis of each patient’s tumor, not only on the individual variant level, but also on the global variant-landscape level. This comprehensive method of analysis provides multi-dimensional insight on pathways affected in the context of drug-gene or drug-variant interactions.
Call us
855.225.4068Location
303 Wyman Street, Office #349
Waltham, MA 02451
